The recipe
cw.copy = existing.CytoscapeWindow (), it copies the graph back to R from Cytoscape

The code is available here. It is excerpted and explained below. But first, a note about the structure of this -- and other -- demo scripts you will find on this website. They usually contain one function, 'run', which takes a 'levels' argument. The typical use is to execute the script one step at a time:
run (0)
run (1)
...
run (7)
run (0:7)
Global variables are used in the script, so that values from a previous step are still defined at the subsequent step. In my presentation here, I will just give the body of each run level, each step. Look at the source to see this in context.
Step 0: create the graph
nRows <<- 10
adMatrix <<- matrix (round (runif (nRows * nRows)), ncol=nRows)
adMatrix <<- adMatrix * upper.tri (adMatrix, diag=T)
colnames (adMatrix) <<- 1:nRows
g <<- new ("graphAM", adjMat = adMatrix, edgemode="directed")
Step 1: initialize the variables which will hold three node and three edge attribute lists
nodeList <<- nodes (g)
nodeListLength <<- length (nodeList)
from <<- c ()
to <<- c ()
nodeInteger <<- c ()
nodeChar <<- c ()
nodeFloat <<- c()
edgeInteger <<- c ()
edgeChar <<- c ()
edgeFloat <<- c ()
Step 2: generate the node and edge attributes
for (i in seq (1, nodeListLength)) {
node = nodeList [i]
nodeInteger <<- c (nodeInteger, round (runif (1, max = 500)))
chars = paste (sample (letters), collapse="")
nodeChar <<- c (nodeChar, chars)
nodeFloat <<- c (nodeFloat, runif (1, max = 500))
for (j in seq (i, nodeListLength)) {
node2 = nodeList [j]
if (adMatrix [i, j] == 1) {
from <<- c (from, node)
to <<- c (to, node2)
edgeInteger <<- c (edgeInteger, round (runif (1, max = 500)))
chars <<- paste (sample (letters), collapse="")
edgeChar <<- c (edgeChar, chars)
edgeFloat <<- c (edgeFloat, runif (1, max = 500))
} # if
} # for j
} # for i
Step 3: add the node attributes to the graph
g <<- initNodeAttribute (g, "nodeInteger", "integer", 0)
nodeData (g, nodes(g), "nodeInteger") <<- nodeInteger
g <<- initNodeAttribute (g, "nodeChar", "char", '')
nodeData (g, nodes(g), "nodeChar") <<- nodeChar
g <<- initNodeAttribute (g, "nodeFloat", "numeric", 0.0)
nodeData (g, nodes(g), "nodeFloat") <<- nodeFloat
Step 4: add the edge attributes to the graph
g <<- initEdgeAttribute (g, "edgeInteger", "integer", 0)
edgeData (g, from, to, "edgeInteger") <<- edgeInteger
g <<- initEdgeAttribute (g, "edgeChar", "char", '')
edgeData (g, from, to, "edgeChar") <<- edgeChar
g <<- initEdgeAttribute (g, "edgeFloat", "numeric", 0.0)
edgeData (g, from, to, "edgeFloat") <<- edgeFloat
Step 5: create the window, then display and layout the graph
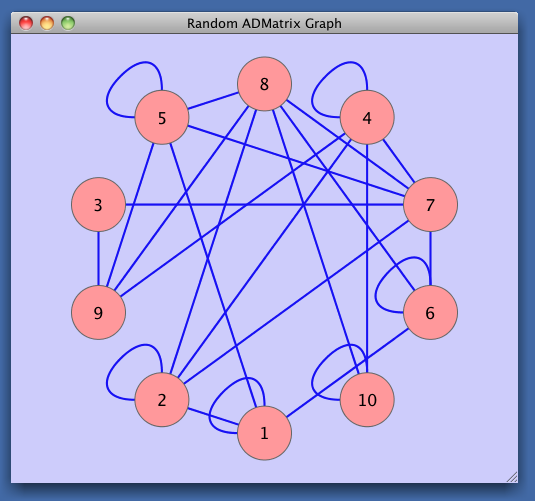
window.name <<- 'Random ADMatrix Graph'
if (window.name %in% as.character (getWindowList (cy)))
destroyWindow (cy, window.name)
cw <<- CytoscapeWindow (window.name, g)
displayGraph (cw)
layoutNetwork (cw, "jgraph-circle")
redraw (cw)
Step 6: Retrieve a copy of the graph from Cytoscape
cw2 <<- existing.CytoscapeWindow (window.name, copy.graph.from.cytoscape.to.R=TRUE)
Step 7: check for equality between the original graph -- nodes, edges, and attributes -- and the one returned
checkEquals (sort (nodes (cw@graph)), sort (nodes (cw2@graph)))
checkEquals (sort (edgeNames (cw@graph)),sort (edgeNames (cw2@graph)))
original.edge.attribute.names <<- sort (eda.names (cw@graph))
retrieved.edge.attribute.names <<- sort (eda.names (cw2@graph))
added.attributes <<- c ('interaction', 'canonicalName', 'edgeType') # Cytoscape adds the first two attributes to all edges; RCytoscape the third.
checkEquals (sort (c (added.attributes, original.edge.attribute.names)), retrieved.edge.attribute.names)
edgeNames.with.bars = sort (names (edgeData (cw@graph))) # for instance, "1|1" "1|2" "1|5" ...
edgeNames.tokenized = strsplit (names (edgeData (cw@graph)), '\\|')
edjat.names = eda.names (cw@graph) # obtain the edge attribute names, used in the inner loop below
for (name.pair in edgeNames.tokenized) {
a = name.pair [1]
b = name.pair [2]
for (ea in edjat.names) {
g1.value = edgeData (cw@graph, a, b, ea)
g2.value = edgeData (cw2@graph, a, b, ea)
checkEquals (g1.value, g2.value)
} # for ea
} # for name.pair